...
Place the jar file in the desired installation directory. The remainder of this document assumes that it is located in in directory $PHYLONET_PATH/jar. Installation is now complete. In order to run PhyloNet, you must execute the file PhyloNet_X.Y.Z.jar, as described in the next section.
...
| Code Block |
|---|
>java -jar $PHYLONET_PATH/PhyloNet_X.Y.Z.jar script.nex
|
Where $PHYLONET_PATH is the path directory of jar file PhyloNet_X.Y.Z.jar.Where , and script.nex is the NEXUS file containing the commands to be executed.
...
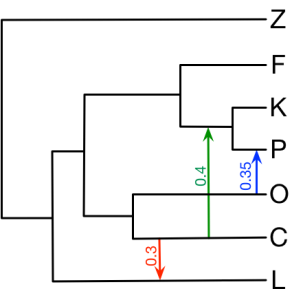
Here we provide all the input nexus files in section 4 of the book chapter. The figure below is the true network we would like to infer. You can also find all the commands below on https://wiki.rice.edu/confluence/display/PHYLONET/List+of+PhyloNet+Commands .
4.1 Minimize deep coalescent Inference
...
4.1.1 MDC Inference using true gene tree topologies
| Input nexus NEXUS file | Maximum number of reticulations |
|---|---|
| InferNetwork_MP_pl8_0_true.nex | 0 |
| InferNetwork_MP_pl8_1_true.nex | 1 |
| InferNetwork_MP_pl8_2_true.nex | 2 |
| InferNetwork_MP_pl8_3_true.nex | 3 |
| InferNetwork_MP_pl8_4_true.nex | 4 |
...
4.1.2 MDC Inference using gene tree topologies estimated by IQTREE
| Input nexus NEXUS file | Maximum number of reticulations |
|---|---|
| InferNetwork_MP_pl8_0_false.nex | 0 |
| InferNetwork_MP_pl8_1_false.nex | 1 |
| InferNetwork_MP_pl8_2_false.nex | 2 |
| InferNetwork_MP_pl8_3_false.nex | 3 |
| InferNetwork_MP_pl8_4_false.nex | 4 |
...
4.2.1 ML Inference using true gene tree topologies
| Input nexus NEXUS file | Maximum number of reticulations |
|---|---|
| InferNetwork_ML_pl8_0_true.nex | 0 |
| InferNetwork_ML_pl8_1_true.nex | 1 |
| InferNetwork_ML_pl8_2_true.nex | 2 |
| InferNetwork_ML_pl8_3_true.nex | 3 |
| InferNetwork_ML_pl8_4_true.nex | 4 |
...
4.2.2 ML Inference using gene tree topologies estimated by IQTREE
| Input nexus NEXUS file | Maximum number of reticulations |
|---|---|
| InferNetwork_ML_pl8_0_false.nex | 0 |
| InferNetwork_ML_pl8_1_false.nex | 1 |
| InferNetwork_ML_pl8_2_false.nex | 2 |
| InferNetwork_ML_pl8_3_false.nex | 3 |
| InferNetwork_ML_pl8_4_false.nex | 4 |
...
4.2.3 ML Inference using true gene tree topologies and branch lengths
| Input nexus NEXUS file | Maximum number of reticulations |
|---|---|
| InferNetwork_ML_bl_pl8_0_true.nex | 0 |
| InferNetwork_ML_bl_pl8_1_true.nex | 1 |
| InferNetwork_ML_bl_pl8_2_true.nex | 2 |
| InferNetwork_ML_bl_pl8_3_true.nex | 3 |
| InferNetwork_ML_bl_pl8_4_true.nex | 4 |
...
4.2.4 ML Inference using gene tree topologies and branch lengths estimated by IQTREE
| Input nexus NEXUS file | Maximum number of reticulations |
|---|---|
| InferNetwork_ML_bl_pl8_0_false.nex | 0 |
| InferNetwork_ML_bl_pl8_1_false.nex | 1 |
| InferNetwork_ML_bl_pl8_2_false.nex | 2 |
| InferNetwork_ML_bl_pl8_3_false.nex | 3 |
| InferNetwork_ML_bl_pl8_4_false.nex | 4 |
...
4.3.1 MPL Inference using true gene tree topologies
| Input nexus NEXUS file | Maximum number of reticulations |
|---|---|
| InferNetwork_MPL_pl8_0_true.nex | 0 |
| InferNetwork_MPL_pl8_1_true.nex | 1 |
| InferNetwork_MPL_pl8_2_true.nex | 2 |
| InferNetwork_MPL_pl8_3_true.nex | 3 |
| InferNetwork_MPL_pl8_4_true.nex | 4 |
...
4.3.2 MPL Inference using gene tree topologies estimated by IQTREE
| Input nexus NEXUS file | Maximum number of reticulations |
|---|---|
| InferNetwork_MPL_pl8_0_false.nex | 0 |
| InferNetwork_MPL_pl8_1_false.nex | 1 |
| InferNetwork_MPL_pl8_2_false.nex | 2 |
| InferNetwork_MPL_pl8_3_false.nex | 3 |
| InferNetwork_MPL_pl8_4_false.nex | 4 |
...
4.3.3 MPL Inference using bi-allelic marker data
| Input nexus NEXUS file | Maximum number of reticulations |
|---|---|
| 0 | |
| 1 | |
| 2 | |
| 3 | |
| 4 |
...
4.4.1 MCMC_SEQ: Bayesian inference on the sequence alignment data
| Input nexus NEXUS file | Maximum number of reticulations |
|---|---|
| 0 | |
| 1 | |
| 2 | |
| 3 | |
| 4 |
...
4.4.2.1 MCMC_GT sampling using true gene tree topologies
| Input nexus NEXUS file | Maximum number of reticulations |
|---|---|
| MCMC_GT_pl8_0_true.nex | 0 |
| MCMC_GT_pl8_1_true.nex | 1 |
| MCMC_GT_pl8_2_true.nex | 2 |
| MCMC_GT_pl8_3_true.nex | 3 |
| MCMC_GT_pl8_4_true.nex | 4 |
...
4.4.2.2 MCMC_GT sampling using gene tree topologies estimated by IQTREE
| Input nexus NEXUS file | Maximum number of reticulations |
|---|---|
| MCMC_GT_pl8_0_false.nex | 0 |
| MCMC_GT_pl8_1_false.nex | 1 |
| MCMC_GT_pl8_2_false.nex | 2 |
| MCMC_GT_pl8_3_false.nex | 3 |
| MCMC_GT_pl8_4_false.nex | 4 |
...
4.4.3 MCMC_BiMarkers: Bayesian inference on the bi-allelic markers
| Input nexus NEXUS file | Maximum number of reticulations |
|---|---|
| 0 | |
| 1 | |
| 2 | |
| 3 | |
| 4 |
...